 |
| Recombinant DNA technology result |
Recombinant DNA technology makes use of science’s understanding of the molecular structure of DNA, the nucleic acid that encodes genetic information, to alter DNA in order to manipulate genetic traits. Such technology has immense implications for agriculture, horticulture, and the generation of medicinal compounds from plants.
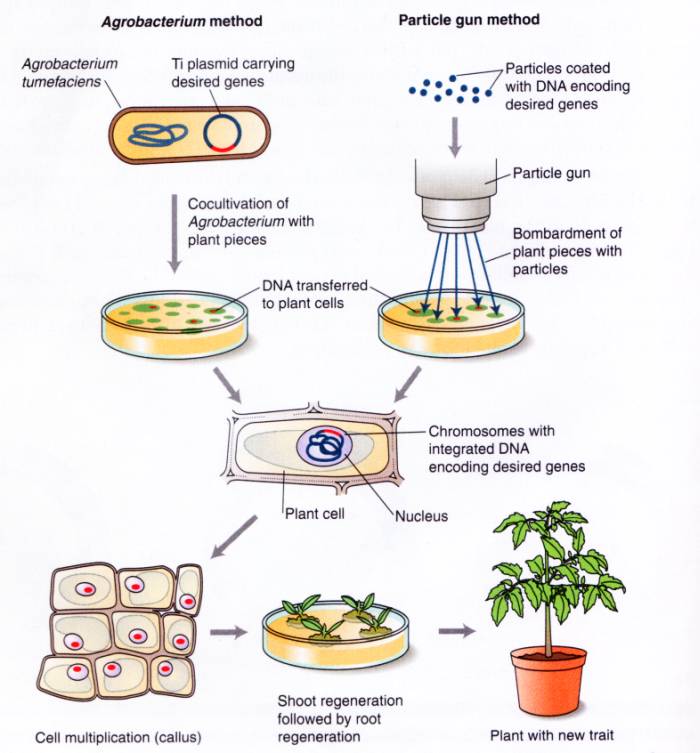
Recombinant DNA technology has been essential for understanding DNA sequences. Because of their large, complex genomes, it was difficult to study one gene in eukaryotes, but recombinant DNA technology has allowed the isolation and amplification of specific DNA fragments facilitating the molecular analysis of genes. In addition, the tools of recombinant DNA technology have been used to create genetically modified plants.
Such modifications include the introduction of resistance to insects, herbicides, viruses, and bacterial and fungal diseases into plants. Plants have also been made to produce antibodies so that plants can serve as edible vaccines.
  |
DNA Structure
Organisms contain two kinds of nucleic acids: ribonucleic acid, or RNA, and deoxyribonucleic acid, or DNA. DNA is made of a double chain, or helix. The structure of one chain, or strand, is a backbone made up of repetitions of the same basic unit.
That unit is a five-carbon sugar molecule called 2′-deoxyribose attached to a phosphate residue. RNA contains a ribose sugar instead.Also attached to the sugar part of the backbone are other molecules called bases. The four bases are adenine (A), guanine (G), cytosine (C), and thymine (T).
DNA molecules are double strands that are held together because each base in one strand is paired to (hydrogen-bonds with) a base in the other strand. Adenine always pairs with the base thymine, and guanine always pairs with cytosine. A and T are called complementary bases. Likewise, Gand Care complementary bases.
DNA is shaped much like a helical ladder, with the sugar and phosphate backbones being the sides of a ladder and the base pairs that hold the two strands together being the rungs of a ladder.
DNA is often represented as a string of letters, with each letter representing a base. The order of A’s, T’s, G’s, and C’s (the rungs of the ladder) along a DNA double helix is the sequence of that DNA which contains the genetic information.
Restriction Enzymes
 |
| Recombinant plasmid with restriction enzymes |
In recombinant DNA technology, scientists are able to use molecules called restriction enzymes to make cuts at specific sequences. Some restriction enzymes make cuts straight across the two strands of DNA in the double helix, creating blunt ends. Other restriction enzymes cut the two strands in a staggered pattern, leaving short, specific single strands at the cut sites.
These single-stranded regions, called sticky ends or cohesive ends, can base-pair (hydrogen-bond) with complementary base sequences from other, similarly cut DNAs. These sticky ends allow joining of DNA from any source cut with restriction enzymes that create the same ends.
Cutting with restriction enzymes creates fragments of DNA with sequence-specific ends that can be spliced into small, self-replicating vehicle, or vector, molecules and introduced into a host cell where the vector molecules with the added DNA fragments replicate to produce a large amount of specific DNA for analyses. This process is recombinant DNA cloning.
   |
DNA Cloning
One way to clone a specific gene is to clone all the DNA fragments generated from cutting with a restriction enzyme and then screen for the clone containing the desired gene. This method of cloning random DNA segments into a vector is called shotgunning.
The entire collection of such cloned fragments, which together represent the entire genome of the organism, is called a gene library. Genomic DNA libraries are made by cloning the total genomic DNA of an organism.
Another way to clone a specific gene is to begin with messenger RNA (mRNA) from the organism. (Messenger RNA is a molecule that functions to create complementary copies of DNA strands.
At a ribosome the messenger RNA then determine the order of amino acids that are joined to make a protein.) Using reverse transcriptase (an enzyme encoded by some RNA viruses that uses RNA as a template for DNAsynthesis), scientists can make a DNA copy of the mRNA.
 |
| DNA cloning |
The complementary strand is also synthesized to create a double-stranded DNA called cDNA (complementary DNA) that is complementary to the mRNA. These cDNAs are then cloned to create a complementary DNA library. Individual cloned cDNAs can be used to trap the corresponding mRNA on a nitrocellulose filter.
At this point, the mRNA can be used in a cell-free protein synthesis system to allow identification of the protein encoded by that cDNA clone. Alternatively, the cDNA can be used to find sequences complementary to it in a genomic library to obtain a clone of the specific gene.
Nucleic Acid Hybridization
The ability to hybridize nucleic acids to find sequences complementary to a particular DNA is another essential tool that offers another way to identify cloned genes. This method is called nucleic acid hybridization or Southern blotting (named after E.M. Southern,who developed themethod).
In this procedure, DNA is cut with restriction endonucleases, and the resulting DNA fragments are separated by size using agarose gel electrophoresis. The DNA in the gel is denatured (made single-stranded) by high pH and transferred to a nitrocellulose filter.
The DNAs are immobilized on the nitrocellulose in the same pattern as on the gel (a Southern blot). A probe—a specific DNA or RNA—is hybridized to the nitrocellulose. The probe is “labeled” with a radioactive or fluorescent (nonradioactive) tag so it can be detected.
The probe is denatured by heat so it is single-stranded and able to anneal (hybridize) with its complementary sequence among the single-stranded DNAs tethered to the nitrocellulose. The probe is then detected to reveal the position of the DNAs that hybridized with the probe.
Polymerase Chain Reactions
Another tool of molecular biology is to use a polymerase chain reaction (PCR) to amplify specific segments of DNA in vitro (in the test tube). PCR requires a pair of sequences, called primers, about twenty base pairs long that are complementary to the ends of the region of DNA to be amplified.
High temperature is used to denature the double-stranded DNA. At a lower temperature, the primers anneal (base-pair) to their complementary sequences, and a thermal-stable DNA polymerase copies the single-stranded templates.
After the replication of the segment between the two primers (one cycle), the newly synthesized double-stranded DNA molecules are denatured by high temperature, the temperature is lowered, primers anneal, and a second cycle of replication occurs. The number of DNA molecules produced doubles with each cycle of replication.
As a result, a million copies of a single DNA molecule can be produced in only a few hours, if the appropriate sequences for the two primers are known. PCR is a very sensitive method: Even a single DNA molecule can be amplified. PCR is much faster than recombinant DNA cloning and can produce a large amount of a specific piece of DNA.
Developing ways to determine DNA sequences (the sequences of adenine, thymine, guanine, and cytosine base pairs on the DNA “rungs”) has led to the identification of the complete DNA sequences of the genomes of a number of organisms, including the model plant Arabidopsis as well as the much more complex human genome.