 |
| Chromatin |
Chromatin is an inclusive term referring to DNA and the proteins that bind to it, located in the nuclei of eukaryotic cells. The huge quantity of DNA present in each cell must be organized and highly condensed in order to fit into the discrete units of genetic material known as chromosomes. Gene expression can be regulated by the nature and extent of this DNA packaging in the chromosome, and errors in the packaging process can lead to genetic disease.
Scientists have known for many years that the hereditary information within plants and other organisms is encrypted in molecules of deoxyribonucleic acid (DNA) that are themselves organized into discrete hereditary units called genes and that these genes are organized into larger subcellular structures called chromosomes.
James Watson and Francis Crick elucidated the basic chemical structure of the DNA molecule in 1952, and much has been learned since that time concerning its replication and expression.
  |
At the molecular level, DNA is composed of two parallel chains of building blocks called nucleotides, and these chains are coiled around a central axis to form the well-known double helix.
Each nucleotide on each chain attracts and pairs with a complementary nucleotide on the opposite chain, so a DNA molecule can be described as consisting of a certain number of these nucleotide base pairs.
The entire human genome consists of more than six billion base pairs of DNA, which, if completely unraveled, would extend for more than 2 meters. It is a remarkable feat of engineering that in each human cell this much DNA is condensed, compacted, and tightly packaged into chromosomes within a nucleus that is less than 10–5 meters in diameter.
Plants typically have larger genomes than humans; for example, wheat has fifteen billion base pairs of DNA. By contrast, the most widely studied plant among current scientists is Arabidopsis. The species Arabidopsis thaliana was selected as a model organism in plant research because of its comparatively simple structure: Its 26,000 genes make up “only” 125 million base pairs.
 |
| Zoomed DNA |
What is even more astounding is the frequency and fidelity with which this DNA must be condensed and relaxed, packaged and unpackaged, for replication and expression in each individual cell at the appropriate time and place during both development and adult life. The essential processes of DNA replication or gene expression (transcription) cannot occur unless the DNA is in an open or relaxed configuration.
Chemical analysis of mammalian chromosomes reveals that they consist of DNA and two distinct classes of proteins, known as histone and nonhistone proteins. This nucleoprotein complex is called chromatin, and each chromosome consists of one linear, unbroken, double-stranded DNA molecule that is surrounded in predictable ways by these histone and nonhistone proteins.
The histones are relatively small, basic proteins (having a net positive charge), and their function is to bind directly to the negatively charged DNA molecule in the chromosome.
Five major varieties of histone proteins are found in chromosomes, and these are known as H1, H2A, H2B, H3, and H4. Chromatin contains about equal amounts of histones and DNA, and the amount and proportion of histone proteins are constant from cell to cell in all higher organisms, including the higher plants.
In fact, the histones as a class are among the most highly conserved of all known proteins. For example, for histone H3, which is a protein consisting of 135 amino acid “building blocks,” there is only a single amino acid difference in the protein found in sea urchins as compared with the one found in cattle.
This is compelling evidence that histones play the same essential role in chromatin packaging in all higher organisms and that evolution has been quite intolerant of even minor sequence variations between vastly different species.
Nonhistones as a class of proteins are much more heterogeneous than the histones. They are usually acidic (carrying a net negative charge), so they will most readily attract and bind with the positively charged histones rather than the negatively charged DNA.
Each cell has many different kinds of nonhistone proteins, some of which play a structural role in chromosome organization and some of which are more directly involved with the regulation of gene expression. Weight for weight, there is often as much nonhistone protein present in chromatin as histone protein and DNA combined.
Nucleosomes and Solenoids
The fundamental structural subunit of chromatin is an association of DNA and histone proteins called a nucleosome. First discovered in the 1970’s, each nucleosome consists of a core of eight histone proteins: two each of the histones H2A, H2B, H3, and H4.
Around this histone octamer is wound 146 base pairs of DNA in one and three-quarters turns (approximately eighty base pairs per turn). The overall shape of each nucleosome is similar to that of a lemon or a football.
 |
| Nucleosomes |
Each nucleosome is separated from its adjacent neighbor by about fifty-five base pairs of linker DNA, so that in its most unraveled state they appear under the electron microscope to look like tiny beads on a string. Portions of each core histone protein protrude outside the wound DNA and interact with the DNA that links adjacent nucleosomes.
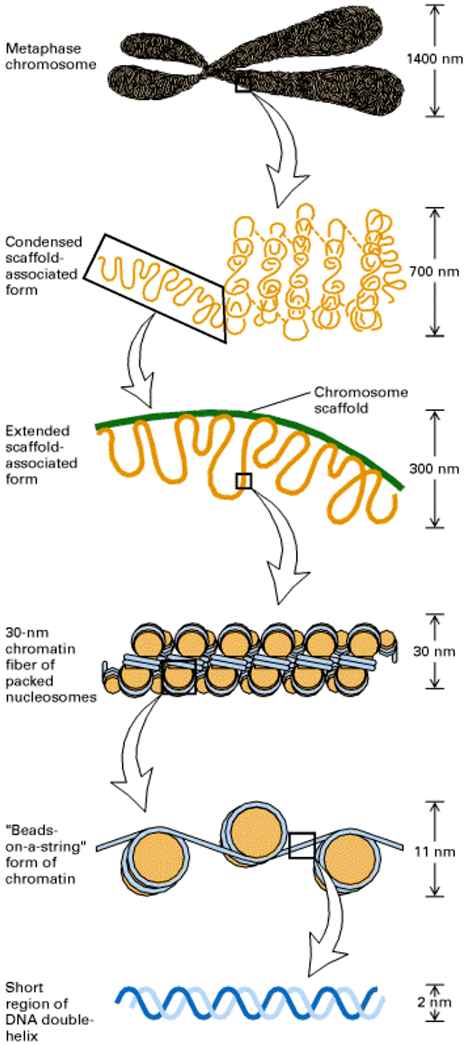
The next level of chromatin packaging involves a coiling and stacking of nucleosomes into a ribbonlike arrangement, which is twisted to form a chromatin fiber about 30 nanometers in diameter, commonly called a solenoid.
Formation of solenoid fibers requires the interaction of histone H1, which binds to the linker DNA between nucleosomes. Each turn of the chromatin fiber contains about twelve hundred base pairs (six nucleosomes), and the DNA has now been compacted by about a factor of fifty.
The coiled solenoid fiber is organized into large domains of 40,000 to 100,000 base pairs, and these domains are separated by attached nonhistone proteins that serve both to organize and to control their packaging and unpackaging.
Loops and Scaffolding
Physical studies using the techniques of X-ray crystallography and neutron diffraction have suggested that solenoid fibers may be further organized into giant supercoiled loops.
The extent of this additional looping, coiling, and stacking of solenoid fibers varies, depending on the cell cycle. The most relaxed and extended chromosomes are found at interphase, the period of time between cell divisions.
Interphase chromosomes typically have a diameter of about 300 nanometers. Chromosomes that are getting ready to divide (metaphase chromosomes) have the most highly condensed chromatin, and these structures may have a diameter of up to 700 nanometers.
One major study on the structure of metaphase chromosomes has shown that a skeleton of nonhistone proteins in the shape of the metaphase chromosome remains even after all of the histone proteins and the DNA have been removed by enzymatic digestion. If the DNA is not digested, it remains in long loops (10 to 90 kilobase pairs) anchored to this nonhistone protein scaffolding.
Impact and Applications
Studies on chromatin packaging continue to reveal the details of the precise chromosomal architecture that results from the progressive coiling of the single DNA molecule into increasingly compact structures. Evidence suggests that the regulation of this coiling and packaging within the chromosome has a significant effect on the properties of the genes themselves.
In fact, errors in DNA packaging can lead to inappropriate gene expression and developmental abnormalities. In humans, the blood disease thalassemia, several neuromuscular diseases, and even male sex determination can all be explained by the altered assembly of chromosomal structures.
The unifying lesson to be learned from these examples of DNA packaging and disease is that DNA sequencing studies and the construction of genetic maps will not by themselves provide all the answers to questions concerning genetic variation and genetic disease.
An understanding of genetics at the molecular level depends not only on the primary DNA sequence but also on the three dimensional organization of that DNA within the chromosome. Compelling genetic and biochemical evidence has left no doubt that the packaging process is an essential component of regulated gene expression.